The content of this page has not been vetted since shifting away from MediaWiki. If you’d like to help, check out the how to help guide!
The ToAST plugin (Tool for Automated Sporozoite Tracking)
ToAST is a Fiji plugin performing tracking and analysis of motile fluorescent objects highly customized to study malaria sporozoite motility. It is based on a popular MTrack2 plugin for ImageJ additionally performing classification of tracks of objects into circular clockwise (CW) or counter-clockwise (CCW) movement, non-motile (“Attached”) and strongly rotationally moving (“Waving”) objects based on short time intervals. Although it was designed to classify the motility patterns of malaria sporozoites that isolated to a glass slide essentially glide in circles, it may be worthwhile trying the plugin on other types of moving objects especially when addressing the questions of motility directionality.
Usage
The input for ToAST is a binary image, to create one please go to Image › Adjust › Threshold, select the level, press “apply”. You might need to invert the image to make the background value 255, the object – 0. Use of automated thresolding plugins is also possible.
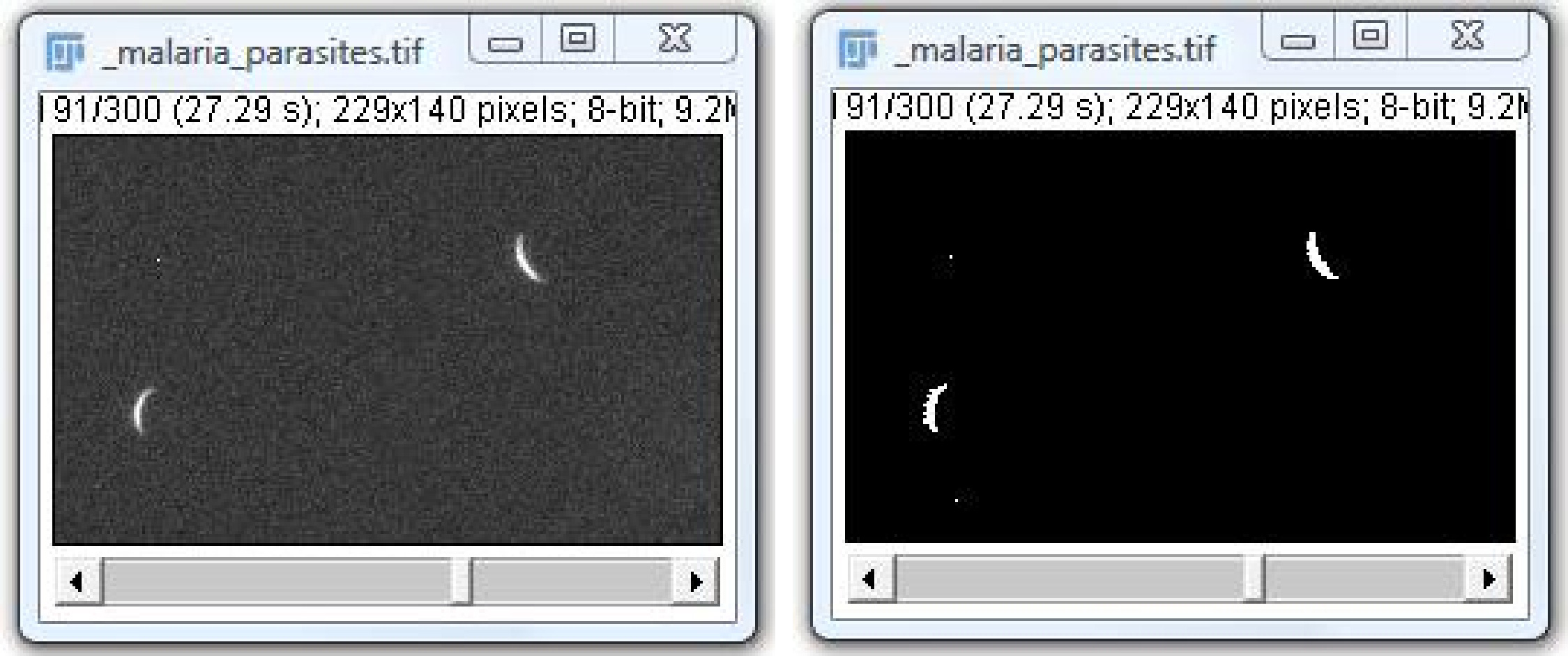
Run Plugins › Tracking › ToAST. In the dialog window please select the parameters of image acquisition.
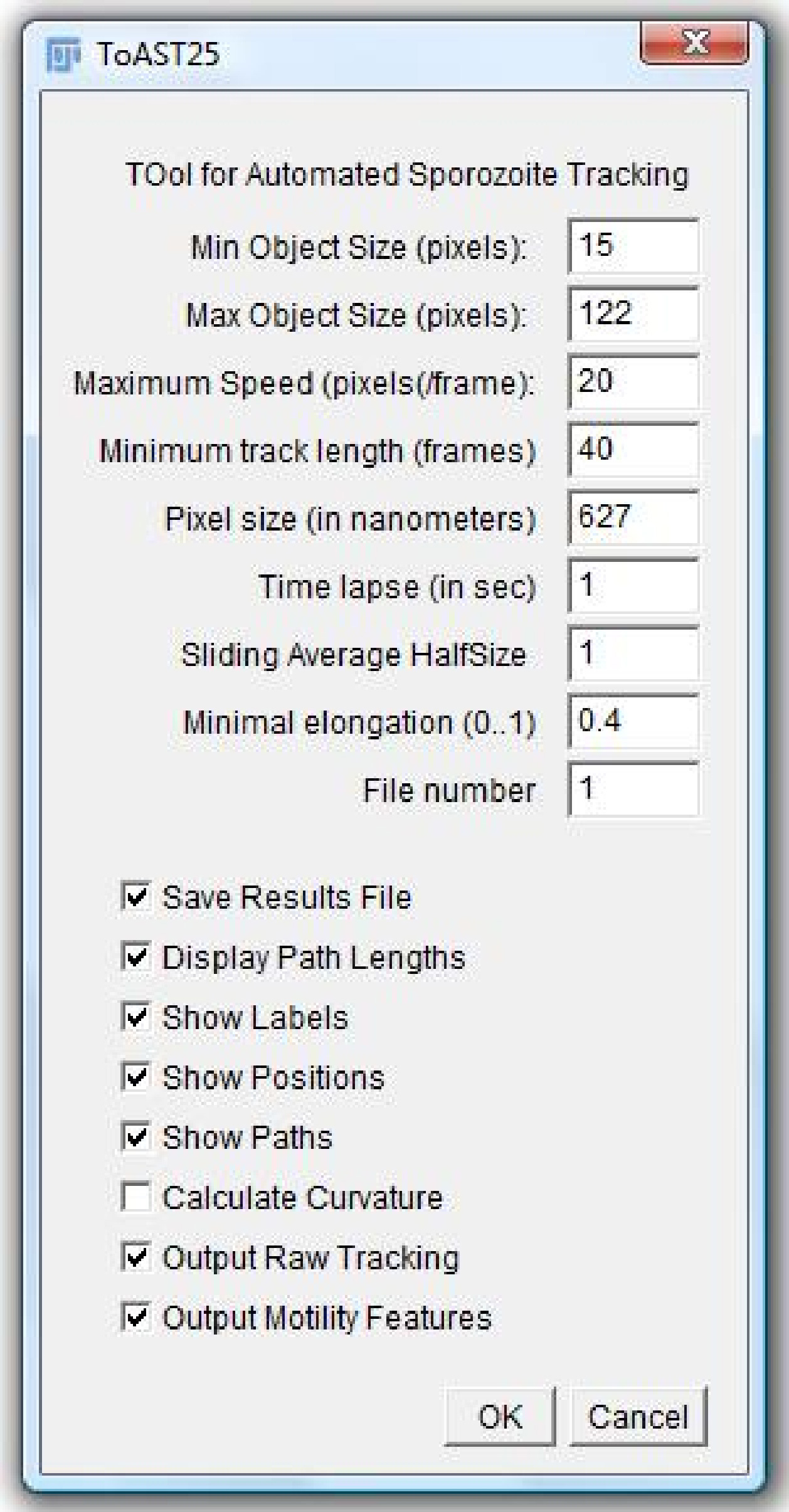
The first 4 parameters are used to distinguish your objects of interest from dirt, overlapping objects and objects that are not quite in focus; it helps to assemble the tracks more reliably. Sliding average half size will serve as a size of the time window (in frames) at which the classification will be performed. All the points that have the speed below 0.5 micrometers per second will be assigned to non-moving; next, all the data points having the rotational speed over 45*translational speed will be assigned to Wavers (high angular velocity and not too high translational); other will be divided into clockwise and counter-clockwise movers.
After pressing OK button the plugin runs creating a log-file where it stores the coordinates, shape factors and motility parameters of the processed movie. It may be imported to any spreadsheet program (such as Excel) by File › Open and then import as space-delimited. First part of the file contains the parameters for every object, every frame.
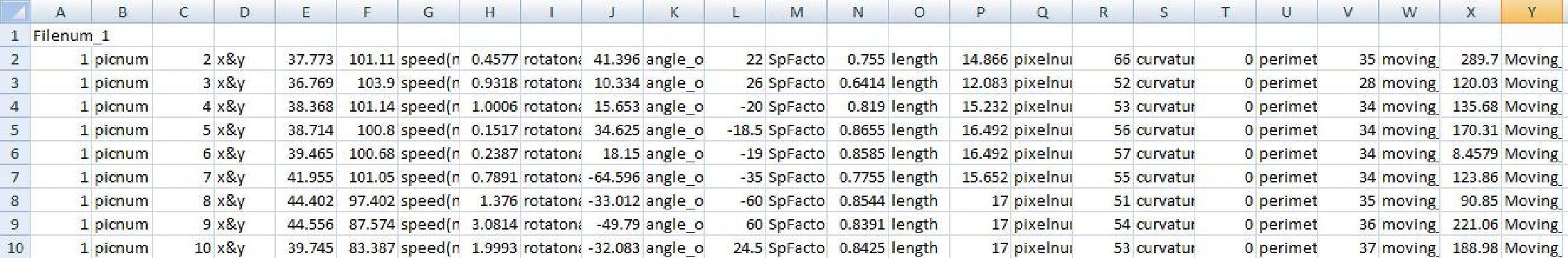
Below you can find all the data again but sorted according to the types of motility; average parameters for every state and frequencies of transmission between states.
Example
There is an example for ToAST in Fiji: File › Open Samples › Malaria Sporozoites (9.2MB)
Citation
- Automated classification of Plasmodium sporozoite movement patterns reveals a shift towards productive motility during salivary gland infection, Hegge S, Kudryashev M, Smith A, Frischknecht F, Biotechnology Journal , 2009, 4, 6 pp: 903-913